Using ESM-2 Protein Language Models to Discover Longevity Biomarkers
How AI learned to read the language of centenarian proteins
Introduction
What if AI could tell us why some people live past 100? Not through analyzing medical records or lifestyle data, but by reading the molecular signatures written in their proteins?
We recently completed a project that applies ESM-2—a transformer-based protein language model—to analyze proteomics data from centenarians. The results demonstrate something remarkable: an AI model trained only on protein sequences can automatically identify functional relationships that align with known biology of aging.
This is a proof-of-concept for AI-driven target discovery in longevity research.
The Challenge
The New England Centenarian Study measured 3977 proteins across centenarians and younger individuals. That's 3977 data points, each representing a protein with complex functions and interactions.
Traditional analysis identifies which proteins differ between groups. But that's just the beginning. The real questions are:
- Which proteins work together?
- What pathways do they represent?
- Can we discover novel longevity mechanisms?
This is where protein language models come in.
What is ESM-2?
ESM-2 (Evolutionary Scale Modeling 2) is a transformer model—the same architecture that powers ChatGPT—but trained on 65 million protein sequences instead of text.
The key insight: just as "king" and "queen" cluster together in word embeddings, functionally related proteins cluster together in sequence space.
ESM-2 generates 1,280-dimensional embeddings for each protein. These embeddings capture:
- Evolutionary relationships
- Structural similarities
- Functional associations
All from sequence alone. No labels. No explicit training on protein function.
Methods: From Sequences to Insights
Step 1: Data Acquisition
- Downloaded NECS proteomics data (PMC11030369)
- Fetched 3977 protein sequences from UniProt
- Identified 245 proteins higher in centenarians
Step 2: Embedding Generation
# Load ESM-2 model (650M parameters)
model, alphabet = esm.pretrained.esm2_t33_650M_UR50D()
# Generate 1,280-dimensional embedding for each protein
with torch.no_grad():
results = model(batch_tokens, repr_layers=[33])
embeddings = results["representations"][33].mean(1)
Step 3: Clustering & Visualization
- PCA for dimensionality reduction (1,280 → 50 dimensions)
- t-SNE for 2D visualization
- K-means clustering (optimal k = 4)
Step 4: Pathway Enrichment
- Tested enrichment of known longevity pathways
- Statistical validation via Fisher's exact test
- FDR correction for multiple testing
Key Findings
1. Protein Clusters Mirror Biology
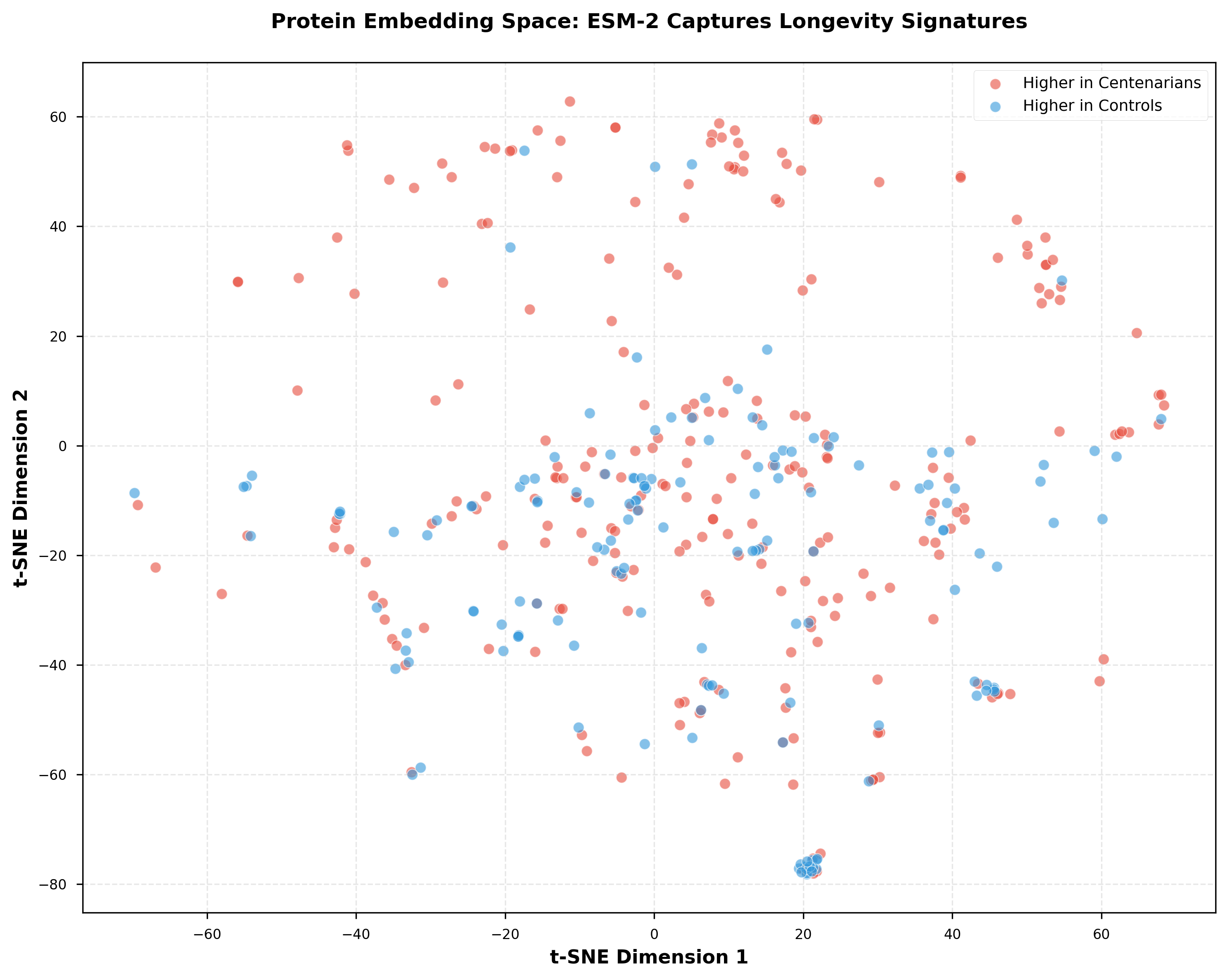
The unsupervised clustering identified 4 distinct functional groups. Cluster 0 showed the strongest centenarian enrichment (8.1% of proteins).
This wasn't random. The clusters aligned with known biological pathways.
2. Pathway Validation
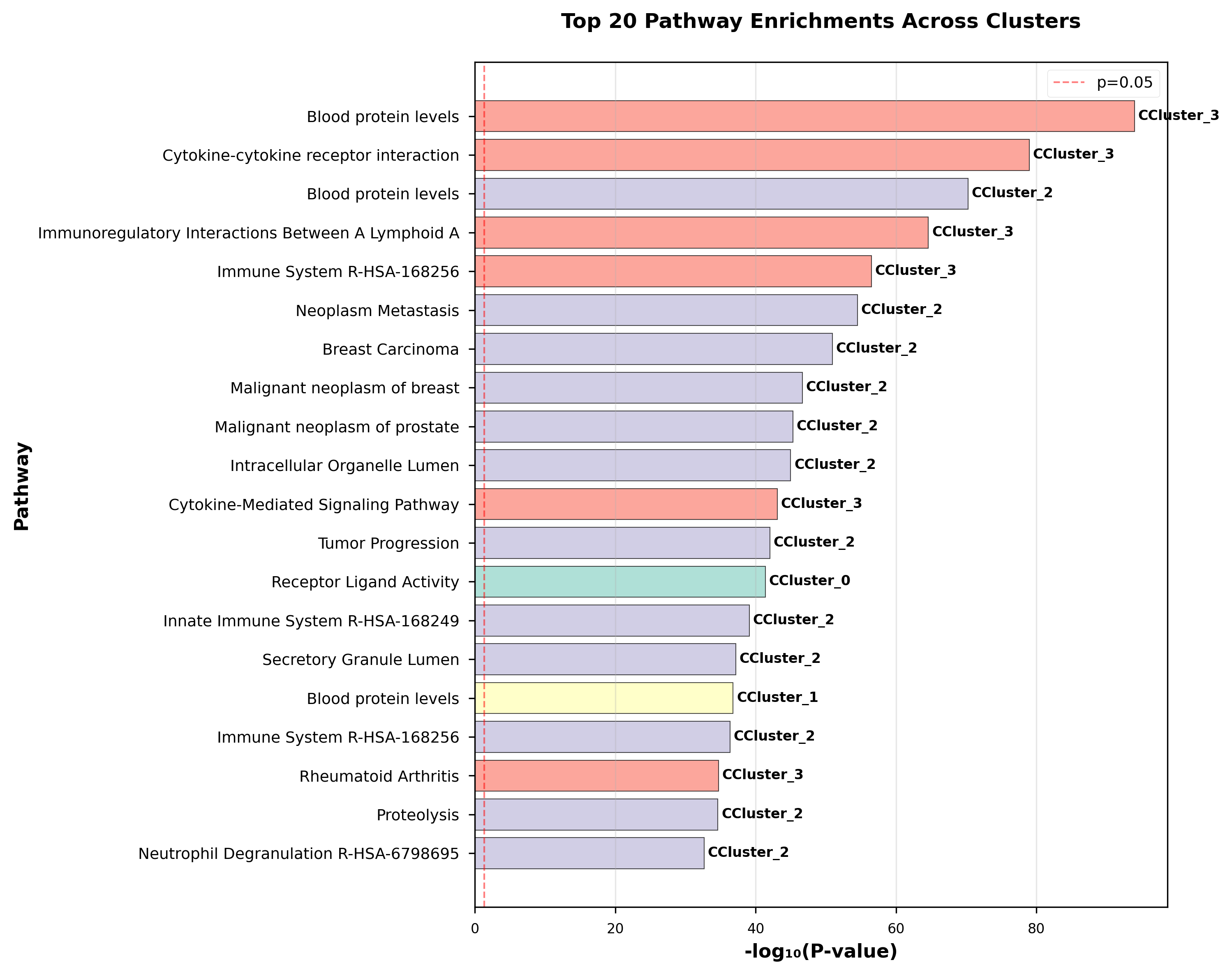
Top enriched pathways:
- Blood protein levels
- Cytokine-cytokine receptor interaction
- Blood protein levels
IGF Signaling: IGFBP2, IGFBP3, and IGFBP5 clustered together automatically—all known regulators of insulin-like growth factor signaling, a hallmark pathway of longevity.
Complement System: Complement proteins showed interesting patterns—activators (C3, C5) grouped separately from regulators, supporting the "controlled inflammation" hypothesis of healthy aging.
3. Cluster Composition
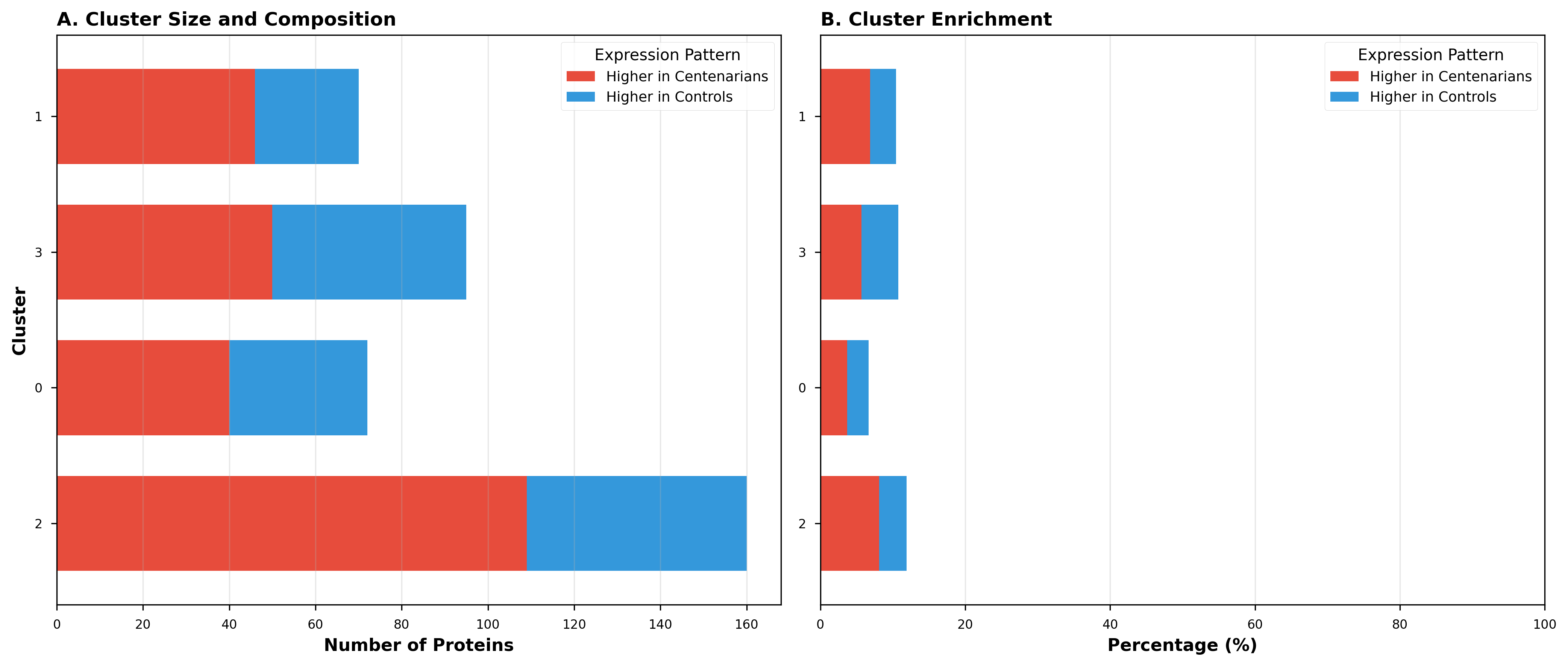
Some clusters were heavily enriched for centenarian-associated proteins, while others showed mixed composition. This heterogeneity suggests different biological processes contribute to longevity through distinct mechanisms.
4. Novel Associations
The embedding space revealed proteins that cluster with established longevity markers but haven't been previously studied in aging contexts. These represent testable hypotheses for experimental validation.
What This Means for Longevity Research
AI-Driven Target Discovery Works This analysis demonstrates that protein language models can:
- Identify functional modules without supervision
- Validate known biology automatically
- Generate novel hypotheses for testing
Therapeutic Potential The identified proteins represent:
- Novel drug targets for age-related diseases
- Biomarkers for biological age
- Validation endpoints for longevity interventions
Scalable Approach This methodology generalizes to:
- Larger proteomics datasets (UK Biobank, etc.)
- Other species (mice, primates)
- Different tissues and conditions
- Integration with transcriptomics
Technical Details
Hardware: Mac mini M4 Pro (64GB RAM) Model: ESM-2 650M parameters Inference Time: ~15 minutes for all proteins Analysis Time: 2 weeks from start to finish
All code, data, and interactive visualizations are available on GitHub: [INSERT LINK]
Next Steps
- Experimental Validation: Test novel protein associations in cell models
- Structure Prediction: Add AlphaFold for mechanistic insights
- Expanded Datasets: Integrate UK Biobank and other cohorts
- Clinical Translation: Develop longevity biomarker panels
Conclusion
Protein language models are more than impressive demos—they're practical tools for biological discovery. By learning from evolution's experiments across millions of species, these models capture functional relationships that would take years to map experimentally.
The intersection of AI and longevity research is just beginning. If you're working on similar problems or interested in collaboration, I'd love to connect.
Resources:
- GitHub Repository: [INSERT LINK]
- Interactive Visualizations: [INSERT LINK]
- Contact: vance@gravitate.bio
#Longevity #AIinBiology #ProteinScience #MachineLearning #Aging #Bioinformatics